Science and Technology has been active in the life sciences for quite some years. The projects range from extraction and processing of unstructured data (texts) to interpretation of life science data. In the next paragraphs we list a few projects.
Automated solutions for information retrieval from large amounts of data
Bioinformatics is hampered by a data deluge. New methods in genetics, proteomics, metabolomics and biochemics in combination are generating data at an unprecedented pace. Combining these data with information contained in a fast growing number of biomedical publications and structured databases requires automated solutions.
One of the possible strategies to combine information from the aforementioned heterogeneous data sources is to use semantic (web) technology. Semantic technology can be used to overcome the differences in naming between the various information sources.
Key aspects
Data science for Life sciences
Semantic concept recognition based on open source indexers
Development of ontologies and thesauri to be used for semantic applications
Creation of semantic triple stores
Development of applications that can exploit these semantic triple stores
Application
In order to make the information from several sources manageable by computer programs, various formalisms have been invented. One formalism that draws quite some attention is the nano-publication. A nano-publication is a logical predicate (a specified relationship between a subject and object, a so-called triple) with contextual information and provenance data. This formalism has been invented to quickly be able to reason over large sets of triples for various tasks. One possibility is to discover semantic indirect connections (paths) between two data items in the triple store. These relationships can be scored on basis of their newness.
Building a large semantic knowledge graph that consists of biomedical concepts with their relationships is essential in many of the knowledge intensive applications in life sciences. Science and Technology has provided tools and expertise to Euretos on extracting these biomedical concepts with their relationships from texts and databases and storing them in a semantic knowledge graph. Nodes in the graph represent biomedical concepts as defined in a broad range of terminologies (medical concepts primarily from the Unified Medical Language System, UniProt, EntrezGene, etc.) with named relationships between concepts mined from over 200 biomedical databases and PubMed. The semantic knowledge graph can be used by different AI applications in the medical field. One application is the discovery of novel drug-disease associations (drug repurposing).
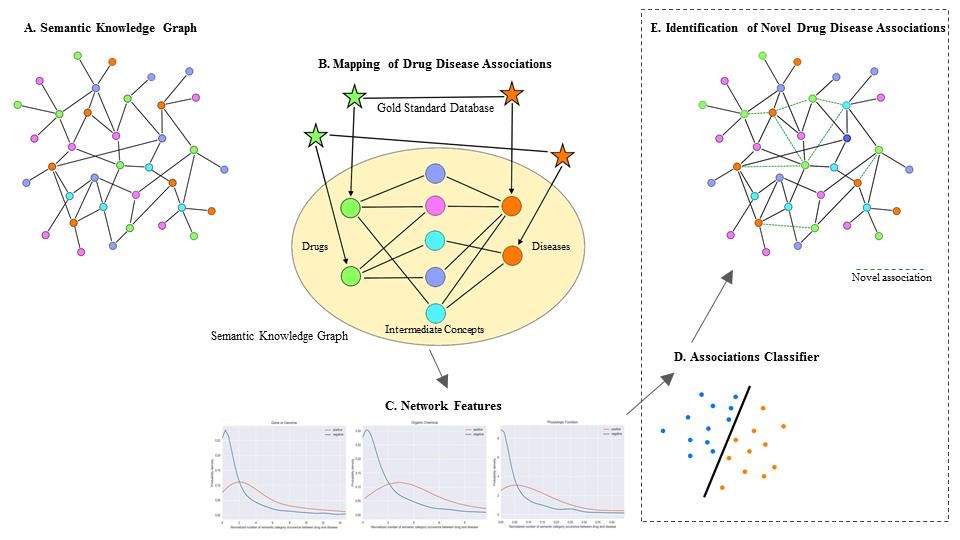
The same semantic knowledge graph has been used to gain insight into the mode of action of anti-inflammatory agents in blood for TNO’s department of microbiology and systems biology. The knowledge graph was extended with specific relations and concepts text mined from documents provided by TNO.
Together with Code-N, a US-based start-up, a number of specific applications around drug repurposing have been developed. Based on information in the semantic knowledge graph new applications for existing drugs were classified and scored.
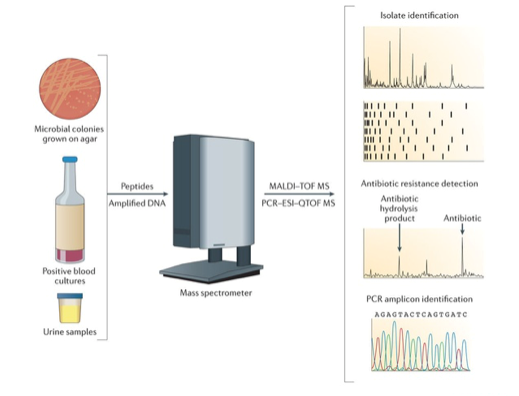
BiosparQ, a spin-off company of TNO, provides a diagnostic platform to classify microorganisms within a few minutes. Based on the mass spectrum the microorganisms contained in the sample can be identified. S[&]T has contributed to the processing of the data from the mass spectrometer and the classification of the spectrum as belonging to a particular microorganism.
For the company ResearchANT S[&]T developed an online experimental compound database. The database is accessible as a web service and can be queried in various ways for experimental compounds. The semantic search has been facilitated by linking the compounds to PubChem’s chemical vocabulary and information contained in PubChem. All compound term variations and identifiers were used to assist the users of ResearchANT in searching for experimental compounds.

Based on large data sets with key-stroke information a machine learning approach has been developed to classify key-stroke patterns as related to a change in user health. S[&]T supported Neurocast (neurocast.nl) with the development of their Neurokeys App. This app uses key-stroke information as a measure of the physical and mental wellbeing of the user. S[&]T developed and implemented machine learning algorithms to classify keystroke patterns to a change in user health.
Life Sciences mobile
Global presence causes geographically dispersed companies, institutes and organizations to have in-house experts across the globe who may never have met in person. In combination with the trend toward expertise specialization, this dispersion creates an inherently difficult quest for the right, available multi-disciplinary team. Keeping track of your organization's expertise can be a "make or break" when competing in a global market. So how do you locate those in-house experts? Searching the company’s intranet for relevant documents? Skimming through internal folder structures trying to estimate their relevance? Asking the company’s senior staff and relying on their typically localized network? In a world of interconnectivity, make the right connection quickly by mining the untapped resource of structured and unstructured data that underlies every organization!
Your untapped data is a goldmine of opportunity!
Most companies have invested the last decades in their information systems: their documents are digital, stored in a document repository system and accessible through the intranet. Information on project execution is stored in a project management system that often offers capabilities to plan and allocate human resources, find optional usage, etc. In many companies, an additional plethora of databases for additional tasks is available: customer-relation databases, manufacturing databases, human resource databases, etc. Apart from the internal data sources a whole range of external data sources is available to find additional information on competitors, the latest developments, etc.
We at S[&]T have developed an expertise analytics platform that automatically constructs an up-to-date expertise profile for employees based on the documents to which they are 'linked'. The S[&]T Expert Platform is an on-premises solution that allows you to locate that one invaluable expert with just the right combination of expertise, experience and availability. Even – or perhaps especially – when you have to find that person from a globally dispersed population of over 100,000 staff. We use machine learning to analyse all your existing documents: intranet/s, CV repositories, reports, presentations, professional social media profiles, and more.
For each employee, the platform creates expertise profiles, allowing you to:
Match CVs with vacancies
Build optimal cross-department project teams
Find a subject-matter expert, within international enterprises
Analyse the impact of mergers, acquisitions, and divestment on expertise position
Track trends over time regarding your expertise position
Define and monitor recruitment targets for their expertise
Visualize expertise assets
Assess competitive landscape from scientifically documented links
Create an intracompany network of specialists or scientists
Ontological Analysis
Ontology: a crucial concept in data mining
An ‘ontology’ is a data structure that describes all relevant search terms as well as the relationships between those entities. The relationships differentiate it from a simple database of entities. For instance, the ontology defines a “narrower-than” relationship between “renewable energy” and “wind energy”. The ontology can be largely created from readily available databases and documents and can also be adjusted to match the client’s domain.
The expertise analytics platform exploits the ontology relationships in the matching process: the search query is automatically expanded with narrower defined concepts; searching for people that have “renewable energy” in their expertise profile would automatically include those people that have “wind energy” in their profile. Candidate concepts not yet covered by the ontology are automatically captured by the term extraction software.
Text is analysed using a suite of semantic tools developed by S[&]T to extract metadata such as named entities (people, organizations), location information (city, country), phone numbers and email addresses. Mentions of known concepts, defined in an ontology, are extracted and so are mentions of new, interesting ontological terms that are not covered by the pre-defined ontology.
March 2019
For more information, please contact Erik van Mulligen.